ChatSpatial MCP Server
MCP server for spatial transcriptomics analysis with 60+ integrated methods
Documentation
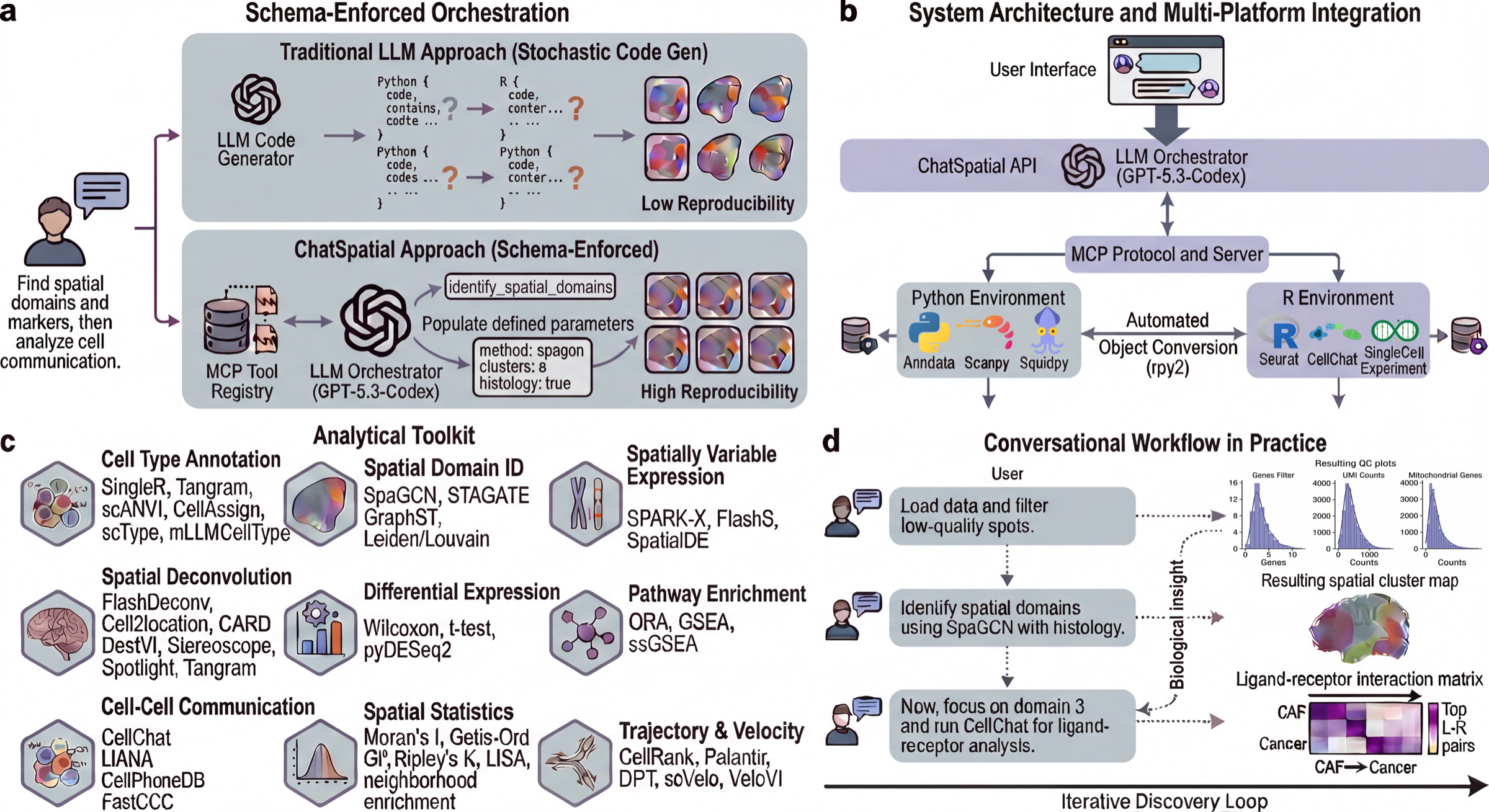
ChatSpatial replaces ad-hoc LLM code generation with schema-enforced orchestration. Instead of generating arbitrary scripts, the LLM selects tools and parameters from a curated registry, making spatial transcriptomics workflows more reproducible across sessions and clients.
ChatSpatial exposes 20 schema-validated MCP tools that orchestrate 65 spatial transcriptomics methods across 15 analytical categories. The tools are the stable natural-language interface; the methods are the analysis backends selected through tool parameters.
Start Here
- Install ChatSpatial — Installation Guide for Python/uv setup, or Docker Guide for the GHCR image
- Configure your MCP client — Configuration Guide
- Run your first analysis — Quick Start
Docker quick start:
docker pull ghcr.io/cafferychen777/chatspatial:v1.2.10
Minimal example prompt:
Load /absolute/path/to/spatial_data.h5ad and show me the tissue structure
If you use Docker, mount host data to /data and prompt with the container path, for example /data/spatial_data.h5ad.
ChatSpatial works with any MCP-compatible client — Claude Code, Claude Desktop, Codex, OpenCode, and other MCP-capable tools.
Capabilities
Current coverage includes 65 methods across 15 analytical categories, exposed through 20 MCP tools. Supports 10x Visium, Xenium, Slide-seq v2, MERFISH, seqFISH.
| Category | Example methods |
|---|---|
| Data Loading & Preprocessing | Scanpy I/O, QC, Normalization, HVG, PCA, Neighbors |
| Visualization | Spatial plots, Embedding plots, Gene expression overlays |
| Spatial Domain Identification | SpaGCN, STAGATE, GraphST, BANKSY, Leiden, Louvain |
| Deconvolution | FlashDeconv, Cell2location, RCTD, DestVI, Stereoscope, SPOTlight, Tangram, CARD |
| Cell-Cell Communication | LIANA+, CellPhoneDB, CellChat (cellchat_r), FastCCC |
| Cell Type Annotation | Tangram, scANVI, CellAssign, mLLMCelltype, scType, SingleR |
| Differential Expression | Wilcoxon, t-test, Logistic Regression, pyDESeq2 |
| Trajectory Inference | CellRank, Palantir, DPT |
| RNA Velocity | scVelo, VeloVI |
| Spatial Statistics | Moran's I, Local Moran, Geary's C, Getis-Ord Gi*, Ripley's K, Co-occurrence, Neighborhood Enrichment, Centrality Scores, Local Join Count, Network Properties |
| Enrichment Analysis | GSEA, ORA, Enrichr, ssGSEA, Spatial EnrichMap |
| Spatially Variable Genes | SpatialDE, SPARK-X, FlashS |
| Multi-sample Integration | Harmony, BBKNN, Scanorama, scVI |
| CNV Analysis | InferCNVPy, Numbat |
| Spatial Registration | PASTE, STalign |
Documentation
| Guide | Use this when... |
|---|---|
| Installation | You need to install ChatSpatial in a Python environment |
| Docker | You want a reproducible container runtime or local dependency resolution fails |
| Configuration | You need exact MCP client syntax or the runtime path model |
| Quick Start | ChatSpatial is installed and you want the first successful analysis |
| Concepts | You need to choose an analysis strategy from a biological question |
| Examples | You want copy-pasteable natural-language workflow prompts |
| Methods Reference | You need canonical tool names, method names, parameters, and defaults |
| Troubleshooting | Setup, data loading, or analysis behavior is not working |
| Full Docs | You want the complete documentation site |
Citation
If you use ChatSpatial in your research, please cite:
@article{Yang2026.02.26.708361,
author = {Yang, Chen and Zhang, Xianyang and Chen, Jun},
title = {ChatSpatial: Schema-Enforced Agentic Orchestration for Reproducible and Cross-Platform Spatial Transcriptomics},
elocation-id = {2026.02.26.708361},
year = {2026},
doi = {10.64898/2026.02.26.708361},
publisher = {Cold Spring Harbor Laboratory},
URL = {https://www.biorxiv.org/content/early/2026/03/01/2026.02.26.708361},
journal = {bioRxiv}
}
ChatSpatial orchestrates many excellent third-party methods. Please also cite the original tools your analysis used.
Contributing
Documentation improvements, bug reports, and new analysis methods are all welcome. See CONTRIBUTING.md.








