ChatSpatial
MCP server for spatial transcriptomics analysis with 60+ integrated methods
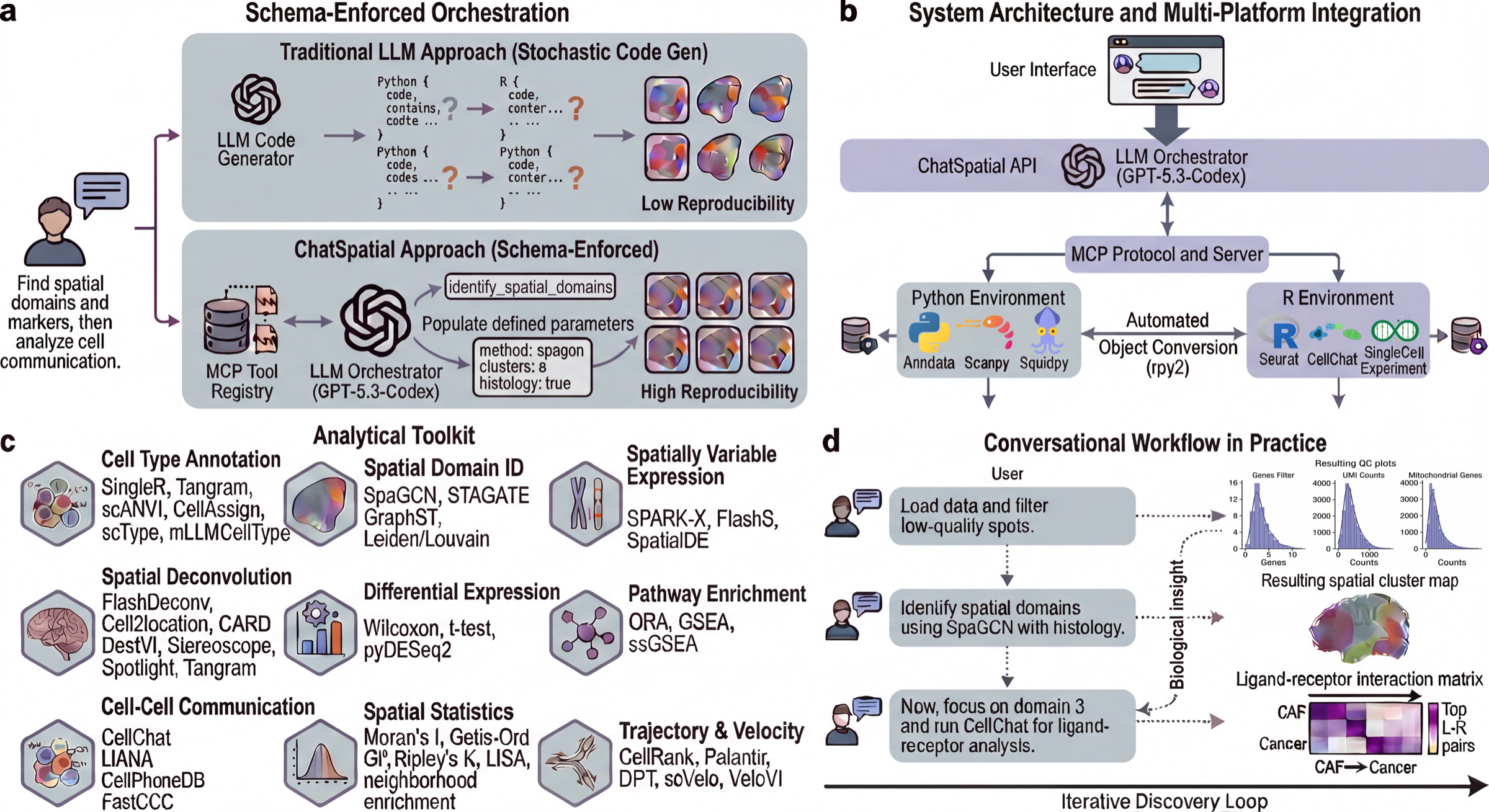
ChatSpatial replaces ad-hoc LLM code generation with schema-enforced orchestration. Instead of generating arbitrary scripts, the LLM selects tools and parameters from a curated registry, making spatial transcriptomics workflows more reproducible across sessions and clients.
It exposes 60+ spatial transcriptomics methods as MCP tools, so any MCP-compatible client can analyze data through natural language.
Start Here
- Install ChatSpatial — Installation Guide for Python/uv setup, or Docker Guide for the GHCR image
- Configure your MCP client — Configuration Guide
- Run your first analysis — Quick Start
Docker quick start:
docker pull ghcr.io/cafferychen777/chatspatial:latest
Minimal example prompt:
Load /absolute/path/to/spatial_data.h5ad and show me the tissue structure
If you use Docker, mount host data to /data and prompt with the container path, for example /data/spatial_data.h5ad.
ChatSpatial works with any MCP-compatible client — Claude Code, Claude Desktop, Codex, OpenCode, and other MCP-capable tools.
Capabilities
60+ methods across 11 categories. Supports 10x Visium, Xenium, Slide-seq v2, MERFISH, seqFISH.
| Category | Methods |
|---|---|
| Spatial Domains | SpaGCN, STAGATE, GraphST, BANKSY, Leiden, Louvain |
| Deconvolution | FlashDeconv, Cell2location, RCTD, DestVI, Stereoscope, SPOTlight, Tangram, CARD |
| Cell Communication | LIANA+, CellPhoneDB, CellChat (cellchat_r), FastCCC |
| Cell Type Annotation | Tangram, scANVI, CellAssign, mLLMCelltype, scType, SingleR |
| Differential Expression | Wilcoxon, t-test, Logistic Regression, pyDESeq2 |
| Trajectory & Velocity | CellRank, Palantir, DPT, scVelo, VeloVI |
| Spatial Statistics | Moran's I, Local Moran, Geary's C, Getis-Ord Gi*, Ripley's K, Co-occurrence, Neighborhood Enrichment, Centrality Scores, Local Join Count, Network Properties |
| Enrichment | GSEA, ORA, Enrichr, ssGSEA, Spatial EnrichMap |
| Spatial Genes | SpatialDE, SPARK-X, FlashS |
| Integration | Harmony, BBKNN, Scanorama, scVI |
| Other | CNV Analysis (InferCNVPy, Numbat), Spatial Registration (PASTE, STalign) |
Documentation
| Guide | Owns |
|---|---|
| Installation | Environment setup, package install, platform notes |
| Docker | GHCR image, volume mounts, and container-backed MCP setup |
| Quick Start | First successful analysis after setup |
| Concepts | Method selection and analysis reasoning |
| Examples | Prompt recipes and workflow examples |
| Configuration | Exact MCP client configuration syntax |
| Troubleshooting | Symptom → fix guidance |
| Methods Reference | Canonical tool parameters and defaults |
| Full Docs | Complete documentation site |
Citation
If you use ChatSpatial in your research, please cite:
@article{Yang2026.02.26.708361,
author = {Yang, Chen and Zhang, Xianyang and Chen, Jun},
title = {ChatSpatial: Schema-Enforced Agentic Orchestration for Reproducible and Cross-Platform Spatial Transcriptomics},
elocation-id = {2026.02.26.708361},
year = {2026},
doi = {10.64898/2026.02.26.708361},
publisher = {Cold Spring Harbor Laboratory},
URL = {https://www.biorxiv.org/content/early/2026/03/01/2026.02.26.708361},
journal = {bioRxiv}
}
ChatSpatial orchestrates many excellent third-party methods. Please also cite the original tools your analysis used.
Contributing
Documentation improvements, bug reports, and new analysis methods are all welcome. See CONTRIBUTING.md.
관련 서버
China Bridge
AI-agent-ready tools for China travel, business, and sourcing. 8 tools including free knowledge guides (payment, visa, VPN), supplier verification, trip concierge, and Stripe ACP checkout.
Cycling Coach AI
AI-powered cycling training coach: personalized training plans, daily workouts, nutrition, strength training, fitness metrics (FTP, VO2Max), and Strava, Garmin, Wahoo, MyWhoosh, and Rouvy integrations.
Immigration & Travel MCP
US visa bulletin data and CBP border wait times. 3 MCP tools for immigration and travel planning.
SwitchBot
Control SwitchBot smart home devices through its official API, enabling automation and integration with AI assistants.
Apviso MCP
MCP server for interacting with the APVISO AI-powered penetration testing platform from Claude Code, Cursor, Windsurf, Codex, and other MCP-compatible tools.
BASTION
Risk Intelligence MCP Server for crypto agents — 52 tools, 72B AI model, 560+ signals, derivatives, on-chain, autonomous trading
Strider Uber Eats
MCP server for Uber Eats food delivery - AI agents can search restaurants, browse menus, and place delivery orders.
Word Orb
The language layer for AI agents. One call returns IPA, definitions, etymology, translations in 47 languages, and ethics guidance — in 3ms.
Weather API MCP Server
Provides current weather data and forecasts using the QWeather API.
Lido MCP Server
Liquid staking data, stETH metrics, and validator info on Lido.








