ChatSpatial
MCP server for spatial transcriptomics analysis with 60+ integrated methods
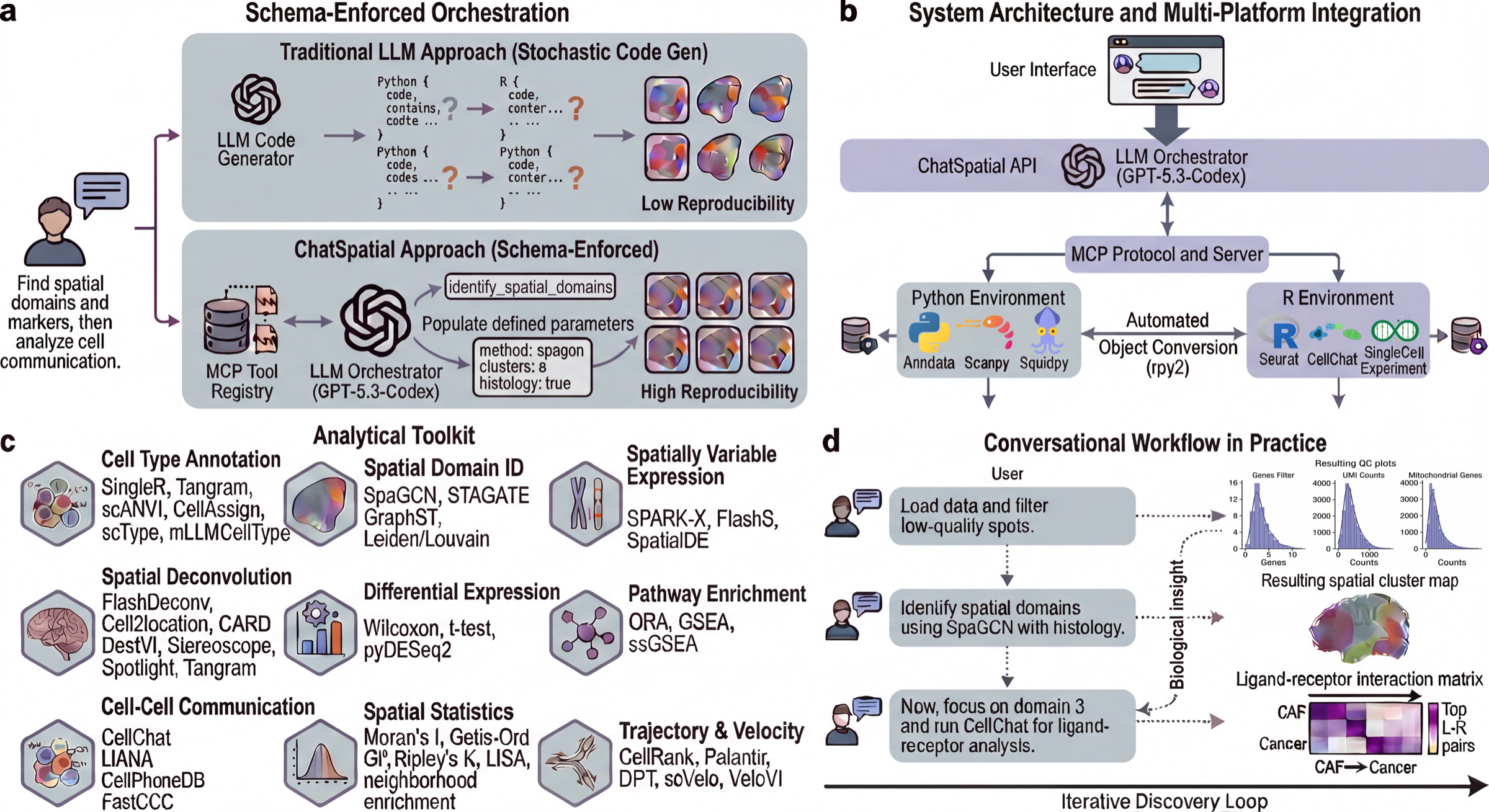
ChatSpatial replaces ad-hoc LLM code generation with schema-enforced orchestration. Instead of generating arbitrary scripts, the LLM selects tools and parameters from a curated registry, making spatial transcriptomics workflows more reproducible across sessions and clients.
It exposes 60+ spatial transcriptomics methods as MCP tools, so any MCP-compatible client can analyze data through natural language.
Start Here
- Install ChatSpatial — Installation Guide for Python/uv setup, or Docker Guide for the GHCR image
- Configure your MCP client — Configuration Guide
- Run your first analysis — Quick Start
Docker quick start:
docker pull ghcr.io/cafferychen777/chatspatial:latest
Minimal example prompt:
Load /absolute/path/to/spatial_data.h5ad and show me the tissue structure
If you use Docker, mount host data to /data and prompt with the container path, for example /data/spatial_data.h5ad.
ChatSpatial works with any MCP-compatible client — Claude Code, Claude Desktop, Codex, OpenCode, and other MCP-capable tools.
Capabilities
60+ methods across 11 categories. Supports 10x Visium, Xenium, Slide-seq v2, MERFISH, seqFISH.
| Category | Methods |
|---|---|
| Spatial Domains | SpaGCN, STAGATE, GraphST, BANKSY, Leiden, Louvain |
| Deconvolution | FlashDeconv, Cell2location, RCTD, DestVI, Stereoscope, SPOTlight, Tangram, CARD |
| Cell Communication | LIANA+, CellPhoneDB, CellChat (cellchat_r), FastCCC |
| Cell Type Annotation | Tangram, scANVI, CellAssign, mLLMCelltype, scType, SingleR |
| Differential Expression | Wilcoxon, t-test, Logistic Regression, pyDESeq2 |
| Trajectory & Velocity | CellRank, Palantir, DPT, scVelo, VeloVI |
| Spatial Statistics | Moran's I, Local Moran, Geary's C, Getis-Ord Gi*, Ripley's K, Co-occurrence, Neighborhood Enrichment, Centrality Scores, Local Join Count, Network Properties |
| Enrichment | GSEA, ORA, Enrichr, ssGSEA, Spatial EnrichMap |
| Spatial Genes | SpatialDE, SPARK-X, FlashS |
| Integration | Harmony, BBKNN, Scanorama, scVI |
| Other | CNV Analysis (InferCNVPy, Numbat), Spatial Registration (PASTE, STalign) |
Documentation
| Guide | Owns |
|---|---|
| Installation | Environment setup, package install, platform notes |
| Docker | GHCR image, volume mounts, and container-backed MCP setup |
| Quick Start | First successful analysis after setup |
| Concepts | Method selection and analysis reasoning |
| Examples | Prompt recipes and workflow examples |
| Configuration | Exact MCP client configuration syntax |
| Troubleshooting | Symptom → fix guidance |
| Methods Reference | Canonical tool parameters and defaults |
| Full Docs | Complete documentation site |
Citation
If you use ChatSpatial in your research, please cite:
@article{Yang2026.02.26.708361,
author = {Yang, Chen and Zhang, Xianyang and Chen, Jun},
title = {ChatSpatial: Schema-Enforced Agentic Orchestration for Reproducible and Cross-Platform Spatial Transcriptomics},
elocation-id = {2026.02.26.708361},
year = {2026},
doi = {10.64898/2026.02.26.708361},
publisher = {Cold Spring Harbor Laboratory},
URL = {https://www.biorxiv.org/content/early/2026/03/01/2026.02.26.708361},
journal = {bioRxiv}
}
ChatSpatial orchestrates many excellent third-party methods. Please also cite the original tools your analysis used.
Contributing
Documentation improvements, bug reports, and new analysis methods are all welcome. See CONTRIBUTING.md.
संबंधित सर्वर
Content Distribution MCP
Multi-platform content distribution - draft, repurpose, schedule, analyze posts for LinkedIn, Instagram, X/Twitter, TikTok. 7 MCP tools.
MCP Kali Server
A comprehensive Model Context Protocol (MCP) server for penetration testing and cybersecurity operations, providing seamless integration between Kali Linux tools and MCP-compatible clients.
AI Dev Jobs
MCP server for the AI Dev Jobs board - 8,400+ open AI/ML engineering roles at 489 companies. Search by role, location, salary, experience level. Live MCP endpoint aidevboard.com/mcp with 4 tools.
CoinGecko MCP Server
Crypto prices, market data, and token metadata via CoinGecko.
MCP Time Server
A simple server that provides the current UTC time.
RateAPI MCP Server
Real interest rates from 1,400+ US credit unions across 50 states. Covers mortgages, auto loans, HELOCs, personal loans, and credit cards. Rates ranked by APR with zero affiliate bias. Works with Claude Desktop and ChatGPT. Free tier available.
Transkribus MCP Server
MCP server for the Transkribus REST API — manage collections, documents, HTR/OCR recognition, models, and more. 290 tools across 22 resource domains.
ROT Trading Intelligence
The first financial intelligence MCP server. Live AI-scored trading signals from Reddit, SEC filings, FDA approvals, Congressional trades, and 15+ sources. 7 tools, 2 resources, hosted remotely, free, no API key required.
N.I.N.A. Advanced API
Control the N.I.N.A. (Nighttime Imaging 'N' Astronomy) software through its Advanced API.
senado-br-mcp
MCP Server for Brazilian Federal Senate open data - legislators, bills, votes, committees








